

Images: Genetic variation of individual kelps helps support the species diversity of the marine ecosystem (left). Meanwhile genetic variation is needed for survival of threatened species, such as the Endangered arroyo oak (right), whose isolated populations are experiencing climate change and drought. Credits: SeacologyNZ.com, The Morton Arboretum
The GEO BON Genetic Composition Working Group, along with collaborators, co-authored a major new and freely accessible publication (Hoban et al. 2022) in the journal “Biological Reviews” that assesses the state of the field for genetic Essential Biodiversity Variables (EBVs). EBVs were established by GEO BON to help aggregate, integrate, and interpret biodiversity observation data from diverse sources, with the ultimate goal of better informing biodiversity conservation. As the authors write, “EBVs measure essential aspects of biodiversity at a higher order than direct observations (e.g., species occurrences), providing a bridge between biodiversity monitoring and decision makers.” The idea is that with aggregated and reliable information and predictive models we can be more certain of biodiversity outcomes in different scenarios, and thus choose the best policy and actions.
Developing genetic EBVs is critical because genetic diversity is the foundation of all biodiversity and supports the stability and resilience of species and ecosystems. Genetic diversity is needed for each species to adapt to challenges, such as climate change- including altered precipitation patterns, extreme weather events, and higher temperatures- and new pests and diseases. Yet it is sometimes ignored in conservation, even though genetic diversity is declining in many species and is poised for precipitous declines in the near future without intervention, as noted in a related paper by members of the Working Group. Thus, Hoban et al. 2022 provides a timely overview of the state of the art of genetic data collection, aggregation, standardization and prediction, which will help inform management actions.
Specifically, Hoban and colleagues present four types of genetic EBVs, evaluate the attributes and feasibility of each as an EBV, document efforts that have been made to aggregate and analyze EBVs from a large number of studies, explain modeling approaches, and discuss data repositories and open data with respect to EBVs. They conclude that the field of genetics is at a critical moment for ensuring common approaches to store, aggregate and analyze the thousands of genetic datasets that already exist.
The four EBVs presented are: genetic diversity, genetic differentiation (the number of populations and the divergence among populations), inbreeding/ relatedness among individuals, and effective population size (a measure of how rapidly genetic diversity is being lost). Together, the four provide a comprehensive picture of genetic composition, but each on its own can provide actionable knowledge. Each EBV has different accessibility, feasibility, costs, data quantities available, and time scale of response to human activities. The authors explain that thousands of scientific studies have recorded genetic EBVs and have “quantified major detrimental anthropogenic drivers [to genetic variation] (e.g., harvesting, selective breeding, habitat change, pollution, reduced population size, isolation, and habitat fragmentation), as well as beneficial anthropogenic drivers such as genetic rescue and migration corridors.” Thus, the genetic EBVs presented will allow us to monitor and report declines in the genetic health of species, and they will help inform appropriate management interventions based on standardized measures.
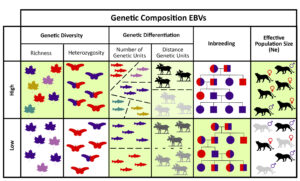
Figure: The four Genetic Composition Essential Biodiversity Variables (EBVs). Green background shading indicates the preferred genetic state (high or low levels) in many conservation or management situations. See Hoban et al for more explanation.
Data accessibility and standardization are essential to ensuring EBVs can be calculated in a distributed and inclusive way. The authors discuss how genetic data are obtained and examined in the laboratory, as well as how genetic data are archived, especially in freely available international online data repositories. However, the authors note that a lack of good metadata (information on the sample date, location, and other attributes) and standard file types and archival databases, means that much of the archived data is not currently usable for large scale analysis of EBVs, as noted by another recent paper by members of the Working Group. Standardization of data collected in the future is vital. In addition, the authors point out the potential of retrospective metadata enrichment, to help rescue older datasets.
The authors describe how EBVs can also help with the re-use of existing genetic data through meta-analyses, mapping projects, and other studies to determine how human activities impact genetic composition. Hoban et al. write that, “overharvested fish populations have 12% lower allelic richness than non-harvested fish,” while plants have “significantly lower genetic richness and evenness in plant populations after habitat fragmentation, especially after more than 100 years of fragmentation.” The paper then discusses the use of modeling environmental drivers for predicting genetic composition EBVs in space and time in order to “project genetic variation into unsampled locations for which data on the driver are available, using the driver as a surrogate of genetic variation” for conservation planning. Genetic-demographic simulations also offer a way to forecast the consequences of specific conservation management actions on genetic EBVs, as explained in an earlier paper by Hoban.
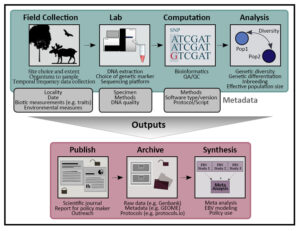
Figure: Steps in generating Genetic Composition Essential Biodiversity Variables (EBVs) include field (or archive) collection of DNA, laboratory work, computational processing of raw data, analysis/calculation, publishing, archiving, modeling and/or synthesis and communication to inform management decisions.
The paper closes with a discussion of challenges and upcoming opportunities. Challenges include the costs of genetic laboratory work, and needs such as “education, training and outreach of practitioners and decision makers”. Among the opportunities are examples of genetic EBVs that are already being used in policy. The paper explains how genetic EBVs have an important role in the European Union Habitats Directive, the US and Canadian endangered species legislation, and in reporting for the post-2020 global biodiversity framework of the Convention on Biological Diversity (CBD), as explained more in another companion paper from the GEO BON Genetic Composition Working Group. The EBV effective population size is currently considered an important indicator for the CBD, while the EBV genetic differentiation could be important for assessing several other indicators. The authors also note that there are interesting connections among EBV “classes” e.g. with the community composition and species populations EBVs already established by GEO BON.
The authors close by advising, “Implementation of our recommendations for data curation, sharing and integration, alongside increasingly detailed assessments of Genetic EBVs, can lead to robust assessment of genetic status and improved conservation and management of the world’s biomes into the future.”
This paper was a collaboration between GEO BON Genetic Composition Working Group, IUCN Conservation Genetics Specialist Group, Society for Conservation Biology Conservation Genetics Working Group, and EU Cost Action Genomic Biodiversity Knowledge for Resilient Ecosystems (G-BiKE), and other participants, such as Genomic Observatories Meta-Database (GEOME).
